All of the biological data is stored in protocols, making it easy to query activity results, filter by condition, and to identify hits via the "Search by protocols" interface on the Explore data tab.
Before setting up your search, make sure that necessary projects are selected in the side-bar: project and data-set scope determines what data will be searched and returned. If nothing is selected, drop-downs will be blank, and no data can be found.
Search across multiple protocols
Filter by readout values, or how to define hit criteria
Search across multiple protocols
The first drop-down box in the "Protocols" search panel of the Explore Data > Search tab allows you to select either “In” or “Not In” to indicate whether the search will return Molecule tested in a Protocol or not yet tested in a Protocol.
The middle drop-down box allows you to specify a Specific Protocol to query, a Protocol Field to query, or a Run Date to query.
- Use the Specific Protocol option to choose a specific Protocol name for your query
- Use Protocol Field to choose a Protocol Field to query (these typically include a Category and Description Protocol-level fields)
- Use Run Date to query across all the Run Dates for all your Protocols
For the Specific Protocol option, the 3rd drop-down box will then allow you to select the specific Protocol to query.
For the Protocol Field option, the 3rd drop-down box will then allow you to search for a keyword in any Protocol Field or select the specific Protocol Field to query.
For the Run Date option, you are now able to use a single query to return all data across all available Protocols using a single Run Date range. (The “earliest” and “latest” query terms are also available.)
You can mix the use of the special earliest/latest terms with the use of actual Run Dates.
This new Run Date querying allows you to find data available across all of your available Protocols during a specified range of dates.
Select a specific protocol
Start by selecting the name of the protocol from the (select protocol) drop-down.
The search is built up sequentially: as you make a selection, more options appear on the page. Once you choose a specific protocol, you can limit your search by exact run date, or by readout definition (result type): all available options will appear under "with run" and "with readout definition" drop-downs.
Quality control search options are available for plate data, and for calculated readouts. You are able to filter out results by error message if a result could not be calculated, or to focus on specific plate data by Z' statistics.
If you are looking for drug selectivity results across efficacy and toxicity assays, or for several quality control parameters in a single assay, multiple protocol terms can be combined into a single search using boolean AND/OR operators.
Filter by run date
Setting this will limit the run dates you are searching.
- (any run): this is the default setting that includes all available run dates in the selected project context.
- within date range: choosing this option brings up a dialog to choose the starting and ending dates for your search. Only available run dates will be populated in the drop-downs.
- with runs in the past: choosing this option brings up a dialog to choose to find Runs over the past 7 or 30 days.
- specific run: choosing this option will bring up a dialog to pick a single date. Runs are listed by date and responsible person. (Additional details may be included in the "Person" field during run creation to make data easier to search).
Filter by condition
If the protocol has any conditions, you will see them listed underneath the "with run" drop-down. For each condition type, there will be a drop-down of available values.

- (any value): this is the default setting that includes all available conditions sets.
- with a specific value for one condition: when a specific condition value is selected, only matching results from the protocol are displayed per molecule.
- specific values for more than one condition: when a specific condition value is selected for each listed condition, returned results will include data that matches ALL set values.
Filter by readout data, or how to define hit criteria
The drop-down in the "any readout definition" section will present all available readout definitions for the selected protocol. If calculated readouts were defined in the protocol, the list will include both the raw readouts and calculated readouts. Subsequent options will depend on the type (raw or calculated), and unit of the readout. Calculated readouts will have additional search options.
Units
There are many units to choose from when defining a readout, but for search purposes, units are broadly categorized into two groups: real numbers and text strings.
- real numbers will include only numeric data, or CDD error messages, and will be searched and ordered according to mathematical rules.
- text strings may be any combination of text and numbers, and will be searched as text only. Mathematical ordering and inequality operators are not applied to text-based readouts even if they contain numeric data.
Operators
Once you choose the readout definition, you can limit results to a subset of values that interest you (hit criteria).
| Operator | Description |
|---|---|
| = | Restrict numeric results to an exact value by entering a number, or to any valid value (not null) by leaving the value blankIC50 = 10 [uM]: Search results will include only exact value 10 IC50 = [uM]: Search results will include all existing IC50 values, including non-numeric values. |
| < | Less than.
IC50 < 10 [uM] |
| > | Greater than.
Inhibition > 95 [%] |
| ≤ | Less than, or equal to.Hill slope ≤ 1 |
| ≥ | Greater than, or equal to.
R squared ≥ .98 |
| from | Use from to search for range of numeric values: from smallest to largest. The search will look for a range ≥ smallest, ≤ largest.% Inhibition from 85 to 102 [%]This search will only return results for tickets that contain both of those words. In other words, both are required. |
|
contains |
Use the contains operator when searching text readouts for a partial match, or a wild-card search. The search will look for any result that matches specified text exactly or as part of another value.Strain contains HL : Returned results include aHL-1, bHL-1, HL-4, HL-9 |
|
does not contain |
Use this operator to exclude results that contain a certain value. Useful when searching text readouts for a partial match, or a wild-card search.
Sequence does not contains SEQGL : Returned sequences do not include SEQGL in any part of the sequence |
| = | For text readouts, this is an exact match operator.
Strain = HELA : Returned results include only HELA |
| ≠ | For text readouts, this is an exact match NOT operator. Returned results will only include values that do not contain entered value
Strain ≠ HELA : Returned results include all strains except HELA. |
Additional search options for calculated readouts
Calculated readouts are created during protocol definition and include Dose-Response calculations and associated readouts or normalized and averaged calculations from raw data. The options appear automatically as a new drop-down list once the appropriate readout is selected in the protocol search panel.
The table below lists only those calculated readouts that are associated with additional search options. If these option do not appear automatically, the selected readout is not a calculation performed by CDD (for example, a final IC50 imported, so while the readout is called IC50, CDD is not performing a data fit and a calculation).
| drop-down option | Readout | Description |
|---|---|---|
| show all | IC50 | This is the default setting, and all molecules with IC50 values will be shown, including all errors. |
| has outliers | IC50 | Show only those molecules. whose IC50 values where the Dose-Response set contains marked outliers. |
| had no error | IC50 | Show only molecules with valid IC50 results, including curves with outliers. |
| has any error | IC50 | Show only molecules with IC50 errors, not including curves with outliers. |
| requires additional points to generate a curve | IC50 | Show molecules whose IC50 could not be calculated due to too few dose-response points (2 points or fewer). |
| could not be calculated | IC50 | Show molecules whose IC50 could not be calculated because no fit could be found for the data points. |
| could not be calculated: insufficient activity | IC50 | Show molecules whose IC50 could not be calculated because the maximum data points did not reach 50% activity. |
| falls outside the dose series | IC50 | Show molecules whose IC50 is either below the lowest tested concentration, or above the highest tested concentration. IC50 results are presented as <lowest conc, or >highest conc. |
Multiple term search
You can add more than one protocol search term by clicking "add a term" underneath the first term. Use this option when you want to look at more than one result from a protocol, or across multiple protocols.

The terms may be related by boolean logic:
AND - all search criteria need to be satisfied in order for the molecule to be returned in the search results.
OR - one or more of the search criteria need to be satisfied in order for the molecule to be returned by the search results.
The terms may be from the same protocol, or a different protocol.
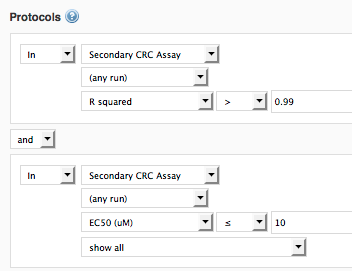
Example 1: Dose-Response Protocol A - find molecules that have curve R2 > 0.99, which have an IC50 value <= 10uM.
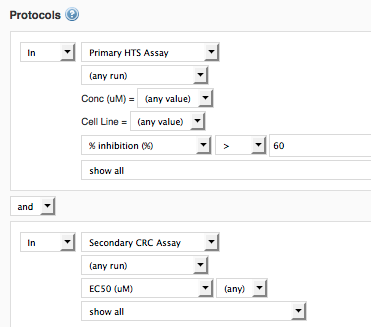
Example 2: Primary SP Protocol X, and Dose-Response Protocol A: find IC50 results of molecules that were hits in the primary screen (Inhibition>60%)
A molecule keyword section is available on the search page as well, and may be combined with the assay data.