Researchers are no strangers to the concept of experimental conditions and parameters, such as cell line, expression type, species, etc, being varied in a single assay. Instead of creating multiple protocols and can be advantageous to create a single protocol which can handle the different conditions simultaneously, while averaging and calculating results based on those conditions. CDD Vault protocols can capture such experimental conditions, and when a protocol readout definition is flagged as a condition, this results in the values of a condition being used to categorize related results for calculated readout definitions within the protocol.
What are protocol conditions
How do you define a protocol condition
How are results aggregated by protocol conditions
How do you import protocol conditions
How do you filter protocol data by conditions
How are results displayed in the search results table
What are protocol conditions
Protocol conditions are a special type of readout definitions in a protocol. The values of a protocol condition are used to categorize related results for aggregation by calculated readout definitions within the protocol. When data are grouped in this way, calculations such as average, median, and IC50 curves, will be performed based on the condition values. For example: An enzymatic concentration-response screen is performed in duplicate with enzymes from 3 different species: Rat, Human and Rhesus monkey, and raw data are imported into CDD Vault for IC50 calculation. The species will be defined as a protocol condition which is applied to the IC50 calculation, resulting in 3 separate IC50 values - based on duplicate data for each species. A protocol may have several such conditions, where each category is based on a combination of condition values.
This can become helpful for later searching and filtering of results in the Explore Data tab as well as how the results are presented in the results table.
How to define a protocol condition
When creating a new or editing any existing readout definition you can "flag" a readout definition with the condition check-box Protocol Condition. Readout definitions of Text, Number, PickList data type are permitted to be conditions, and the flag will appear automatically when one of the two data types is selected.
A best practice is to define conditions as pick-list data type - this data type will ensure the highest data consistency and quality.
You can retroactively mark an existing readout definition as a condition, and the related readout will be re-calculated based on the new condition categories. NOTE: there is a caveat here depending on how many readouts and calculated readout definitions you and how it affects the displayed results. See the section on calculations for more details. Also, to mark an existing readout as a Condition, the readout may not have more than 1000 distinct values across the data already imported into the Protocol.
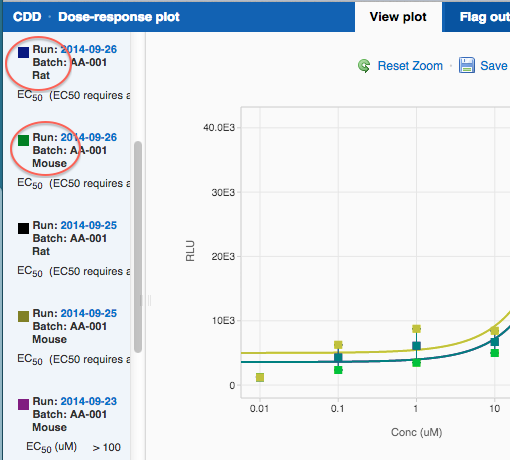
Here's what a IC50 protocol would look like with a species condition defined:

How are results aggregated by protocol conditions
Aggregated readouts to which conditions apply:
- Dose Response calculations: DR series used in e.g. an IC50 calculation will be based on the condition values, so that more than one series may be imported into a single run of a protocol.
- Average calculated readouts based on imported data. When you define an arithmetic mean or geometric mean of a readout, the condition values is taken into account.
- Formula-based calculations available with CDD Vision.
When a protocol condition is defined, aggregated readouts in the protocol will respect the condition values when performing their calculations.
- When a single protocol condition is defined, the grouping will be based on individual values in that condition. To use our Species condition example:
Species= (Mouse, Rat, Rhesus) will result in 3 results - one for each species.
- When more than one protocol condition is defined, the grouping will be based on all possible combinations of values. To use an example of of two conditions "Species" and "Cell Line":
Species = Mouse, Rat, Rhesus
Cell Line = HK-2, MKt-1
resulting in separated calculated readout results for
Mouse, HK-2
Rat, HK-2
Rhesus, HK-2
Mouse, MKt-1
Rat, MKt-1
Rhesus, MKt-1
How do you import protocol conditions
Protocol conditions are imported in the same way as other readout data: condition values need to be present in the import file for each row of data.
NOTE: You will need to make sure that the condition values are spelled correctly, and that a consistent nomenclature is used. CDD Vault will not be able to guess that "rhesus", "Rhezus", and "rhesus monkey", are all the same species, and data will be aggregated incorrectly based on the condition values (hence the recommendation to use pick-lists).
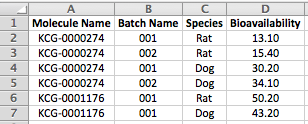
- For single point data, where each row corresponds to a different molecule and batch, the condition values must be listed for each row:
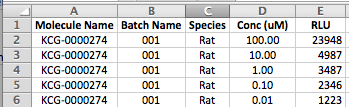
- For concentration-response data, where multiple rows correspond to the same curve per molecule and batch, the condition value must be listed for each row:
- For reference control compounds: specify conditions on each row, just as you would for test compounds.
- For positive and negative controls: no need to specify conditions. Plate controls will be aggregated (averaged) per plate or per run, based on your readout normalization settings.
How do you filter protocol data by conditions
Search for a specific condition value
Since protocol conditions are a special kind of readout, they are displayed separately from other readouts in the Search - Protocols section of the Explore Data tab.
When you select a protocol that contains conditions from the protocols drop-down, the list of condition readouts will be displayed.
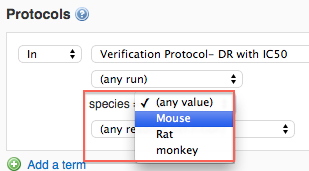
Choose a particular condition value from the drop-down list. The drop-down list will be populated with all available values in your protocol.
How are results displayed in the search results table
When you filter a protocol by a specific protocol condition value, the search results table will return the aggregated readout definition with the standard deviation and number of replicate points (n). The condition value isn't shown in the results table, since you've specifically selected to display data for that one condition value.

On the other hand, when leaving the condition drop-down on "(any value)", you will get results for each aggregated condition value in a separate column.

Please see also the articles on calculations having condition dependencies here:
Custom calculations within one protocol and here: Calculations across multiple protocols.
Only columns that have at least one value for at least one molecule will be displayed automatically. If you want to see a specific set of condition values, set your protocol search terms to filter the data.
For example, for the screenshot above, if you wish to see only human and rat data, compose a query with two terms:
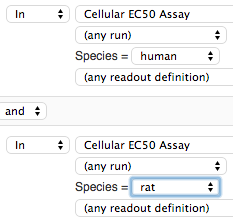
Dose Response IC50's are displayed in the same style as average inhibitions above. Dose-response plots will show individual curves for each condition grouping, and you can see curve legends by opening the separate dose-response window (click on the plot in the search results table).