Files can be attached in several ways. Image data, scans of western blots, lab notebook pages, or analytical data such as NMR spectra, can be securely stored within CDD Vault, associated to the right object. Most objects within the CDD Vault have a "Files" tab where individual files may be attached. Such files are stored on the CDD Vault secure server, and will remain linked with the parent object unless it is deleted. Unless you need store terabytes of files, there is no limitation on the size or file formats that can be attached.
File attachment on the "Files" tab
Files may also be imported as data into molecule/entity or batch fields and protocol readout definitions. This type of file attachment requires fields of the "file" data type to be created ahead of file import. Image files and PDF files will be rendered as preview within the CDD Vault, and may be displayed in-line on the search results page together with the rest of the numeric data.
File attachment on the "Files" tab
Objects within the CDD Vault have a "Files" tab where any file size or file format may be attached for storage and download. However, in the Files subtab of the Molecules and Protocols page you will see a message advising users to use the Molecule and Protocol Fields, respectively, to store the file attachments in these dedicated fields as described below.
Protocols Fields
Attach a protocol methods or SOP to the the protocol record. The file will be stored on the CDD Vault server, and will be available on the "Files" tab of the protocol, or via a "paperclip" icon- shown in the search results table when searching protocol data on the "Explore Data" tab. The icon is a hyperlink that will direct the user to the Protocol details page, and the file attachment may be downloaded from there.
To attach a new file, navigate to the protocol details page by clicking on the protocol name, go to the "Files" tab, and click to "edit attached files". Choose the file on your computer that you wish to attach, and save changes.
Runs
Attach excel spreadsheets of lab notebook pages to the specific runs of protocols. Files attached here are not linked to any specific molecules, so if you need to store readout-level or batch-level file attachments, please read the relevant sections in this article. The file will be stored on the CDD Vault server, and will be available on the "Files" tab of the run, or via a "paperclip" icon- shown in the search results table when searching protocol run data on the "Explore Data" tab. The icon is a hyperlink that will direct the user to the Run details page, and the file attachment may be downloaded from there.
To attach a new file, navigate to the run details page by clicking on the protocol name, then clicking on the run date. Go to the "Files" tab, and click to "edit attached files". Choose the file on your computer that you wish to attach, and save changes.
Molecules
Similarly to protocol and runs, you can attach files to the core molecule/entity record. The file will be stored on the CDD Vault server, and will be available on the "Files" tab of the molecule, or via a "paperclip" icon- shown in the search results table when searching molecules/entities on the "Explore Data" tab. The icon is a hyperlink that will direct the user to the molecule/entity details page, and the file attachment may be downloaded from there.
Navigate to the molecule/entity details record by clicking on the Molecule/Entity name, and go to the "Projects" tab, then follow the steps outlined for protocols.
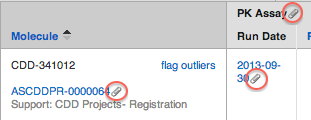
NOTE: As of December, 2020, please store these file attachments in dedicated molecule/entity fields that are created by your vault administrator. CDD Support can help migrate your existing attached files. In the future, once all files previously attached directly to molecules/entities, the molecule/entity attached files tab will be removed.
Files as data
By treating files as a special type of molecule/entity or batch field or readout definition, you can associate images and other files directly with your structures, entities, batches and/or assay data! You can attach any file format or file size in this fashion. CDD Vault will display previews for all common image formats (JPG, GIF, PNG, & BMP) and for PDF files as well. Any other file format will appear as a hyperlink to download the file to your computer.
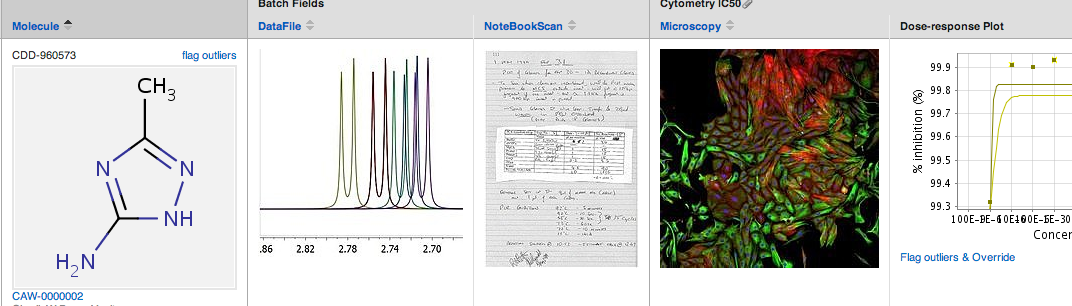
Files attached to protocol readout rows appear in your results table, right alongside the single-point or dose-response data and molecular structure. You can download the original file at any time by clicking on the file name or preview image in any part of the application, for example right within the results table.
Defining File data types
In order to be able to upload files as data into protocols, you will need to add a new readout definition with the "file" data-type. Similarly, in order to add file data to a molecule/entity or batch field, a new field must be defined by the vault administrator with the "file" data-type.
If you're not familiar with setting up protocols and readout-definitions, this should help.
If you're not familiar with molecule/entity and batch fields, this article explains.
For protocol readout definitions:
Navigate to the Protocol details form, and add a new readout definition, making sure to choose the “File” data type. Don't forget to save the changes.
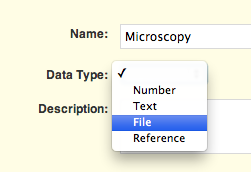
The protocol is now ready to accept files as data either manually or in a bulk upload via the import interface.
For molecule/entity and batch fields:
If you are the vault administrator, go to the "Settings > Vault > Molecule Fields" or "Settings > Vault > Batch Fields" tab, and click to "Add/Edit batch fields". Then define a new field of the "file" data-type. Don't forget to update or save changes.
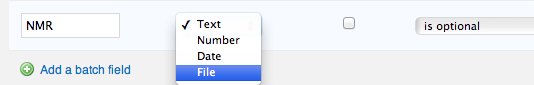
After update, every molecule/entity or batch will now contain a new field, which may be used to attach files manually or in bulk mode.
Attaching files manually
For protocol readout definitions:
If you are adding data manually, you can enter both the numeric and file data at the same time. If you have already uploaded the assay results that are numbers or text, return to the protocol and go to the “All data” tab of the run by navigating to the protocol details page, and clicking on the run date.
On the "All data" tab of the run page, click to edit the desired readout row.
Edit a row and choose a file to attach. You may upload a new file, or choose a previously uploaded file. 
For molecule/entity or batch fields:
Navigate to the Molecule/Entity details tab. Either edit the molecule/entity definition or click on the "Batches" tab and click "edit batch". For any field that was created as type=file, you may upload a new file, or choose from previously uploaded files. 
Bulk file import
For molecule/entity fields, batch fields and protocol readout definitions, multiple files can be attached during a single import by uploading a compressed file (ZIP format) containing all the file attachments, along with the data import file. In order to associate the correct attached file to the correct readout row, the data import file needs to include an additional column or field containing the attached file names.
Create a CSV import file which contains all of the data to be imported:
- Molecule and Batch Identifiers
- Plate and well positions, if needed
- Numeric and text data fields, if needed
- The name of the file you are attaching to each data row: don’t worry if the files are inside a sub-directory, or if you leave off the extensions: CDD Vault will look up the file by name alone, as long as it is not ambiguous.
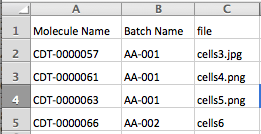
Now create a .ZIP file that contains the following:
- CSV formatted import file
- All of the attachment files- for the example shown above, you would zip together the import file shown in the screenshot, plus files called cells3.jpg, cells4.png, cells5.png and cells6.
On Macs creating a zipped filed is easy. Highlight the files you wish to compress together, right-click, and choose "Compress items". On a PC, you will need to download a free utility such as WinZip.
Upload the compressed file and map the data import file as usual. If the attached file will be imported into a protocol, map it to the “file” readout of the protocol, which you defined earlier (shown in the screenshot below). If the attached file is to be imported into a molecule/entity or a batch field, map it to the corresponding field. If you don't know what I'm talking about, start with the CDD tutorial on data importing.
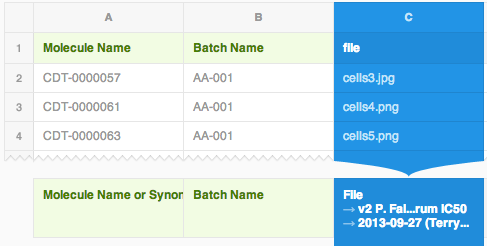
When the import is completed, the attached file previews and download links will be displayed in the detailed forms, as well as search results, and export files.

