Here some details and tips that should help you with curve fitting when the standard calculation doesn’t yield the expected result. Sub-topics are:
- Activity Data Validation
- Z' Factor
- Heatmaps
- Curve Fit Validation
- Data filtering
- Curve Classification
- Plot Inspection and Adjustment
- Outlier flagging
- Curve Fit Parameters Override
- IC50 Override
Activity Data Validation
As soon as you have imported your data, go to the page for that run and look at the Data Summary tab. CDD provides the following tools to detect and investigate problems.
Z′-factor
The Z′-factor (Z prime) describes how well separated the positive and negative controls are, and is widely used as a measure of assay quality. Values between 0.5 and 1 are excellent, values between 0 and 0.5 may be acceptable, and values less than 0 indicate the assay is unlikely to be usable in a high-throughput context. On the run page we display a Z′-factor for each plate, as well as one for the run as a whole. Learn more about plate quality control.
Heat maps
While the Z′-factor conveys assay quality in a single number, heat maps allow you to visualize subtleties in your data and quickly spot problems such as column or edge effects. Click 'View all heat maps for this run' to see all the plates at once. At the drop-down on the left you can select which plate attribute you want to watch in the heat map and set the color scale for that view.
Curve Fit Validation
Most users will want to inspect each dose-response curve individually, and CDD makes it easy to filter and categorize your curves according to pertinent curve fit parameters. A dose-response protocol automatically calculates and provides a comprehensive set of parameters and statistics to quantitatively assess validity of the curve fit in addition to the intercept value.
| Parameter | Description |
|---|---|
| Intercept CI Lower | 95% Lower Confidence limit of the intercept value |
| Intercept CI Upper | 95% Upper Confidence limit of the intercept value |
| Intercept Fold Change | If a reference control is defined on the plate map, the fold change is the ratio of the test compound intercept to the reference compound intercept. Fold change is calculated per plate or per run, depending on the protocol definition. |
| Hill slope | The hill coefficient is one of the 4 parameter equation of the displayed curve fit, as defined in the protocol. |
| Span | Span is defined as the difference between Maximum Response and Baseline Response (in other words, the difference between the two asymptotes of the curve) |
| Baseline Response | The curve minimum or bottom parameter of the 4 parameter equation of the displayed curve fit, as defined in the protocol. |
| Maximum Response | The curve maximum or top parameter of the 4 parameter equation of the displayed curve fit, as defined in the protocol. |
| R-squared | R squared statistic that describes the goodness of curve fit to the actual data points. |
| N | The total number of data points used to perform the curve fit. This includes all of the replicate point. |
| Curve Class | Curve classification score developed by the NIH based on R-squared and maximum measured activity |
| Minimum Measured | Minimum extremal value of all measured responses. NOTE: The minimum activity is not necessarily measured at minimum tested concentration. |
| Maximum Measured | Maximum extremal value of all measured responses. NOTE: The maximum activity is not necessarily measure at maximum tested concentration. |
Data Filtering
From the search page, you can filter the tested compounds based on the curve fit parameters listed in the table above.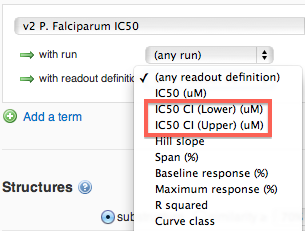
You can also specify an additional criterion on the calculated intersections: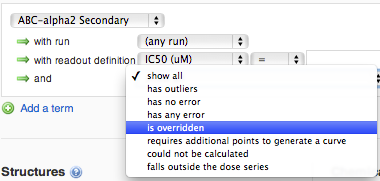
Curve Classification
Please note in order for a curve classification score to be generated, the response variable must be either CDD-normalized data or the plot definition must contain a minimum activity threshold.
One of the available parameters available for validation is "curve class".
Filtering by curve class allows you to easily separate compounds based on the quality of the dose-response curve.
Dose-response curve classification was first proposed in Inglese et al. (Quantitative high-throughput screening: A titration-based approach that efficiently identifies biological activities in large chemical libraries)
Curves are classified based on the quality of curve fit to the data (R2), the magnitude of the response (maximum activity), and the number of asymptotes to the calculated curve.
We have augmented the classification based on conversations with one of the authors of the NCGC CurveFit code by Yuhong Wang, Ajit Jadhav and Noel Southall.
Plot Inspection
Dose-response plots are automatically displayed in the results when you search for a protocol with a dose-response calculation.
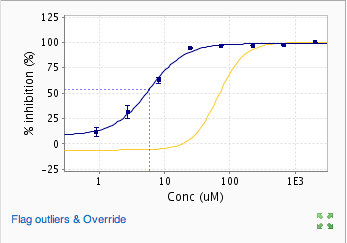
When the fit is successful, it is displayed as well as the sample points and outliers. If you have run replicates, the mean at each dose is plotted with error bars. Error bars represent the standard error of the mean.
Interactive Plot
Click on a plot or the  icon and a new Dose-response window will open. This window can be expanded full screen and the plot resizes accordingly. From there, you can zoom in on any area of interest. Click on the Tips link to learn how to interact with the plot.
icon and a new Dose-response window will open. This window can be expanded full screen and the plot resizes accordingly. From there, you can zoom in on any area of interest. Click on the Tips link to learn how to interact with the plot.
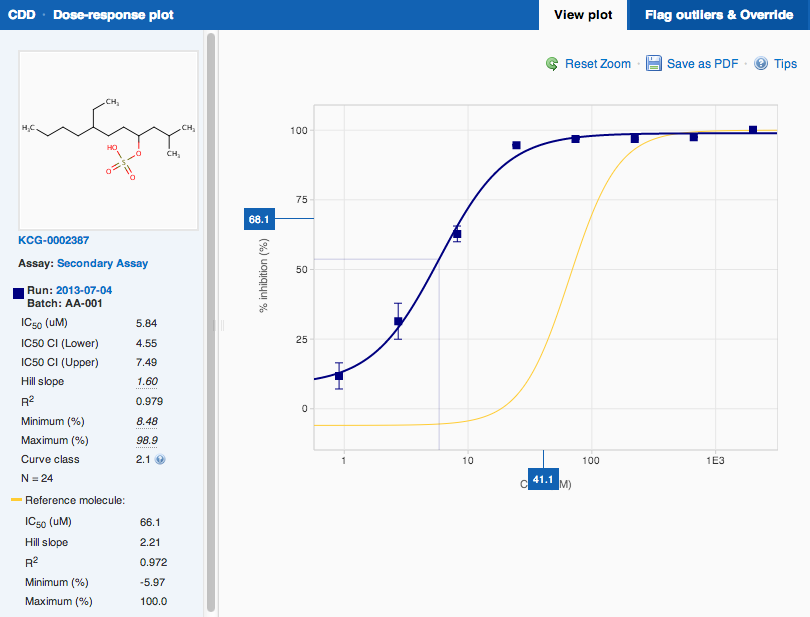
Plot Legend
You can see the plot legend by opening the interactive version of the plot.
Color Coding
When you are visualizing dose-response data on the search results page, the points and curves for a specific run are always drawn using the same color across all plots. The same color is also used for the interactive plot. Outliers and the intercept lines are displayed using a lighter version of that same color.
Reference Substance
Reference substance plot will be overlaid on any sample plot, when displaying data for only one run, if the data you imported includes a reference substance and you have defined it in your control layouts.
When a reference substance is not defined, but a dose-response series is defined for a positive control, this plot will be overlaid on the sample plots.
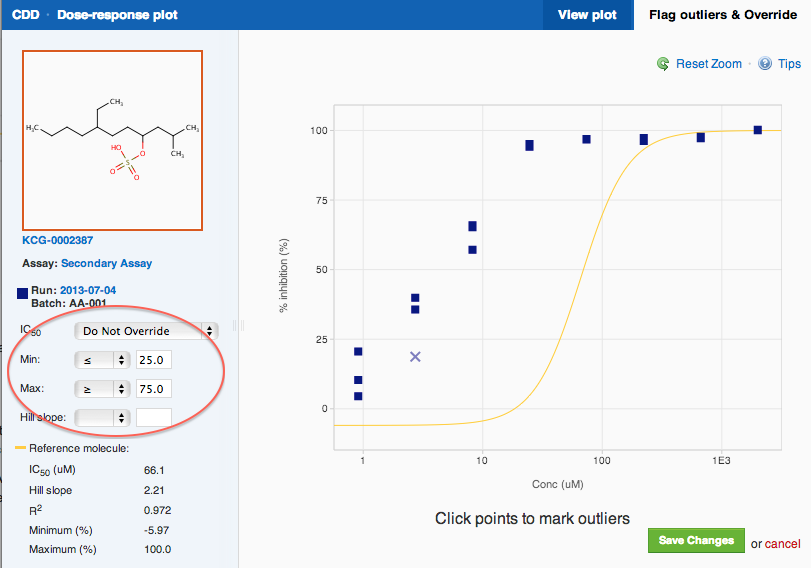
Outlier Flagging
Notice a point that is clearly an outlier? You can flag it as an outlier now through a bulk import as you register new biological data or graphically and textually to data that already lives within your vault.
Graphically
Click on the “Flag outliers and Override” tab in the upper right-hand corner of the window and the Dose-response window opens in edit mode. Zoom in on an area around it and click the point to flag it down. That's it, you are done!
Don't forget to click "Save Changes". The fit and end-point calculations will be adjusted accordingly.
Textually
On the search page, change the display options to include the dose-response protocol activity readouts (raw data, % inhibition, and concentration) then click on the "Flag outliers" link located in the search results table header. Locate the readout and click on the "flag" icon. The result row will refresh with the updated fit data and plot. Note that any calculations that have been affected by overriding parameters or flagging outliers will appear stricken out or underlined in search results.
On the run page, under the “All Data” tab, click on the “Edit” link of the row containing the readout you want to flag as an outlier, then click on the "flag" icon. Save any changes by clicking the “Update this readout” button. The readout row (and any other row for the same batch) will be refreshed with the updated fit.
Bulk Import
Outlier status can now be assigned during a data import. Users can map the outlier status as a separate column, for any numerical readout definition in a protocol. Please note that users do not need to create outlier status as a readout definition in the protocol but rather CDD Vault will automatically give users the option to map an outlier status field for every numerical readout in a specific protocol in the data import wizard. This means that users may conduct a statistical analysis QC for their data prior to upload to identify any outliers in the data set.
Once outliers have been identified, they can be denoted in the upload file in a discrete column by
- “0”, “FALSE”, or simply a blank cell when there is no outlier
- “1” or “TRUE” to indicate when there is an outlier.
In the example below, you can see that a discrete column titled “Efficacy (RLU) Outlier” corresponds to outliers in the “Efficacy (RLU)” numerical readout. The outlier column can then be mapped to the appropriate outlier readout definition. The “1” in the first row indicates this efficacy value will be flagged as an outlier while the second and third rows, blank cells, will not be flagged as outliers.
Once the data is committed, these outliers are flagged and any calculations or aggregations will exclude these values. You can now see that the point we have flagged in the above upload is denoted by an “x” on the dose-response plot below and the IC50 calculated readout is now italicized and underlined indicating that the IC50 value was calculated excluding outliers.
Manual Override of Curve Fit Parameters
If you've already set fit parameters while defining your protocol, you will see that they were applied to the specific curve in the dose-response window. You can choose to override these default parameters either by clearing them completely, which sets them to "float", or by assigning different parameters to this specific curve. The options are the same as for the protocol-level fit parameters, and are found in the Dose-response window you used to mark outliers (see above section). Switch to the "Flag outliers and Override" tab in the top right corner.
Override IC50, EC50, ....
When all else fails, you have the option to override a calculated dose-response value: when the curve fit produces a mathematically correct, but biologically invalid result, and none of the fit adjustments seem to help. This option is located in the Dose-response window you used to mark outliers. Switch to the "Flag outliers and Override" tab in the top right corner, and look in the side-bar. Each curve and each calculated end-point will have their own override. For example, if you calculate an IC90 and an IC99 on the same set of data you will have two overrides.
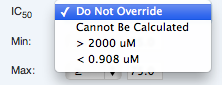
A calculated value may be manually changed to:
- > Max Conc. Tested (e.g. “> 10uM”. This value is determined dynamically for each curve)
- < Min Conc. Tested (This value is determined dynamically for each curve)
- “Cannot be calculated”
- “Do not override” (which restores the automatic calculation)