This article covers the details of dose/concentration response protocol set-up from raw data. We'll assume here that you've created protocols before, so if you have not, you can start with this introductory tutorial instead. Dose-response calculation entails the following steps:
Fit parameters: Min, Max, Hill Slope
Once you've created a new protocol that has, at least, a name and a project, you will need to enter your readout definitions. Generally, numeric readouts (like the Raw Data Measurement and Concentration) will be used in a Dose Response plot/curve, so these numeric readouts will need to be added to your Protocol definition first.
Create a Readout to contain the Raw Data Measurement value:
* Be sure to add the desired normalization setting to this readout!
Create a Readout to contain the Concentration value:
Once these initial readouts (for Raw Data and Concentration) exist, add another Readout definition using the "Plot" data type.
To complete the Dose-Response calculation, fill in the form working from top to bottom.
- start by selecting the "Plot" data type,
- then sect the x and y axes from the drop-down list of existing readouts,
- optionally constrain your fit parameters,
- optionally select a custom Inactive Range, and
- select which data calculation you want (ie, IC50, EC50, etc)
Notes on Normalization
Complete the normalization section of the form even if you don't plan to use CDD for data normalization. Just choose "Do not normalize".
To eliminate plate to plate variation, normalize data by calculating % Inhibition or % Activation based on the positive and negative controls. You'll need to import plate and well data as well as you control data, and we'll go over this here and here. You will also need to assign a name to the normalized readout and choose %inhibition or %activation calculation.
Here's how % Inhibition is calculated: 
Here's how % Activation is calculated: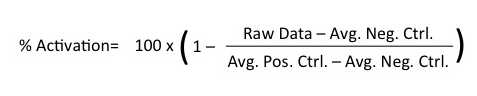
And here's how to define these calculations in your readout definition form:
- Choose a normalization option
- Choose calculation type "% inhibition or activation"
- Name your normalization appropriately
The drop-down for normalization includes the following options which will influence the fit validation as well as plot scales, so choose one that best describes your data:
Normalize within each plate (shown above)- both positive and negative controls are run on each screening plate. Controls will be averaged per plate, and test data normalization will be performed per plate. This will help remove any plate-to-plate variation.
Normalize within each run - positive and negative controls are present on one or on some plates. All controls will be averaged together across plates before test data are normalized.
No controls (do not normalize) - choose this option if you're using raw data that will not have a consistent scale.
Fit parameters: Min, Max, and Hill Slope
The curve fit is performed using the variable Hill slope model or the four parameter logistic curve:
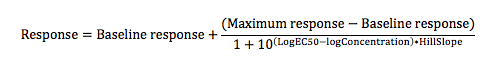
where
Response is the measured response on the Y axis.
Baseline response is the minimum response at the bottom of the plateau.
Maximum response is the maximum response at the top of the plateau.
EC50 is the concentration at 50% response
Concentration is the measured drug concentration on the X axis.
Hill Slope is the Hill coefficient that describes the steepness of the curve.
Using this equation, we perform nonlinear regression with the Levenberg–Marquardt algorithm.
We give you the option to set rules around finding the best fit parameters:
Use best fit - float all parameters, allowing the fit algorithm to automatically fit any value.
Lock fit parameters (shown above)- gives you the ability to define a set of parameters that uniformly applied across all curves in the protocol. The curve fitter will optimize the parameters as usual, except it will enforce the ranges you have specified. You can restrict parameters to a range, but allow them to float within that range. You can define an explicit range (“from”), or a lower (“≥”) or upper (“≤”) bound of each of the three curve fit parameters, in addition to the existing exact number (“=”) option. For example, if you would like all curves in your inhibition assay to have a positive slope, you can set the Hill slope to be ≥ 0. Alternatively, if you wish to fit your dose response curve using the three parameter equation, you will want to set the Hill slope to be = 1, thereby fixing the Hill slope using the standard slope model.
Fit validation: Activity Threshold
You can set the range of activity that defines inactive compounds, so that bogus IC50s will not be calculated for compounds that show insufficient activity. Instead, the reported value for these curves will be “> highest tested concentration”. This setting is part of the dose-response readout definition, and has two options:
At least one point must be OUTSIDE of the set range, in order for CDD Vault to calculate an IC50. If none of the points fall outside the set range, the reported value for these curves will be “> highest tested concentration”
- “3 standard deviations from the negative control mean.” For raw data that is being normalized against controls within CDD Vault, the active compounds may be set to float depending on the imported negative control values.
- “Custom” setting may be used for any curve fit.
Here are a couple of examples for this setting:
Inhibition assay
You are dealing with normalized data, so the range of activity will be from 0% to 100% inhibition. Your compounds should always exhibit inhibitory activity or no activity- there is no chance of activation.
You are measuring IC50.
The activity threshold would be set to < 50%. This means that if all of the measured activity values are < 50%, CDD will report IC50 > highest measured concentration.
Activation assay
You are dealing with normalized data, so the range of activity will be from 0% to 100% activation (you've selected "subtract normalized value from 100%" option when setting up the normalization above) . Your compounds should always exhibit activation activity or no activity- there is no chance of inhibition.
You are measuring EC50.
The activity threshold would be set to < 50%. This means that if all of the measured activity values are < 50%, CDD will report EC50 > highest measured concentration.
Mixed assay: inhibitors and activators
You are dealing with normalized data, but compounds may be either inhibitors (0% to 100% inhibition), or activators (-100% to 0% inhibition) so the range of activity will be from -100% to 100%, and EC50 can be at -50% or 50%
The activity threshold would be set from -50 to 50%. This means that if all of the measured activity values are from -50% to 50%, CDD will report EC50 > highest measured concentration.
Hang in there, you're almost done!
Data Calculations: EC50, IC50, IC99, etc
Select one of the pre-defined end-point calculations, or create your own custom calculation by entering a name and specifying the desired intersection.
For those of you who want more than one calculation for a given data set, we allow you to add additional calculations so you can easily compare without re-importing your data. Are you unsure whether an IC99 or IC98 provides a better approximation of the IC value? Add as many calculations as you want, look at the plots, then remove the ones you don't need. The plots and the calculated data will be automatically updated in the background.
Don't forget to set the display format to the desired number of decimal places or significant figures. This works just like in Excel - the underlying calculations are performed on the full values, but the displayed result is rounded per your display settings.
Now you're done. Don't forget to click "add/update calculation" at the bottom of the form. Here's what the complete form looks like for our example.
Related articles:
